Description
A collection of tools in R to represent and analyse trophic networks in space across aggregation levels. The package contains a layout algorithm specifically designed for trophic networks, using trophic levels and dimension reduction on a diffusion kernel.
Package installation
For the last stable version, use the CRAN version
install.packages("metanetwork")For the development version, use the github install
install_github("MarcOhlmann/metanetwork")Introduction and basics
What is a metanetwork ?
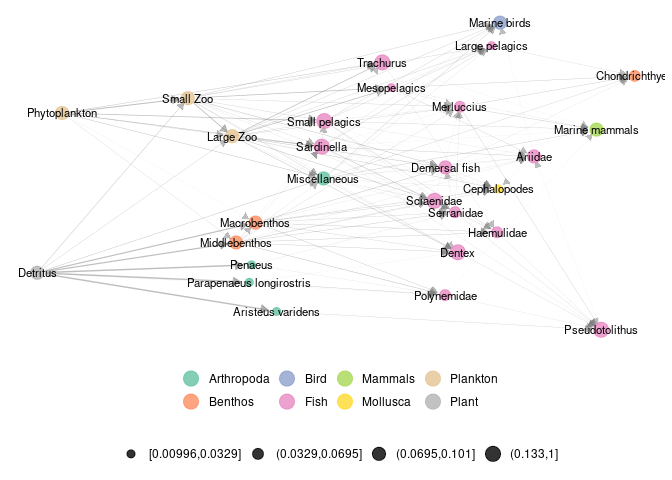
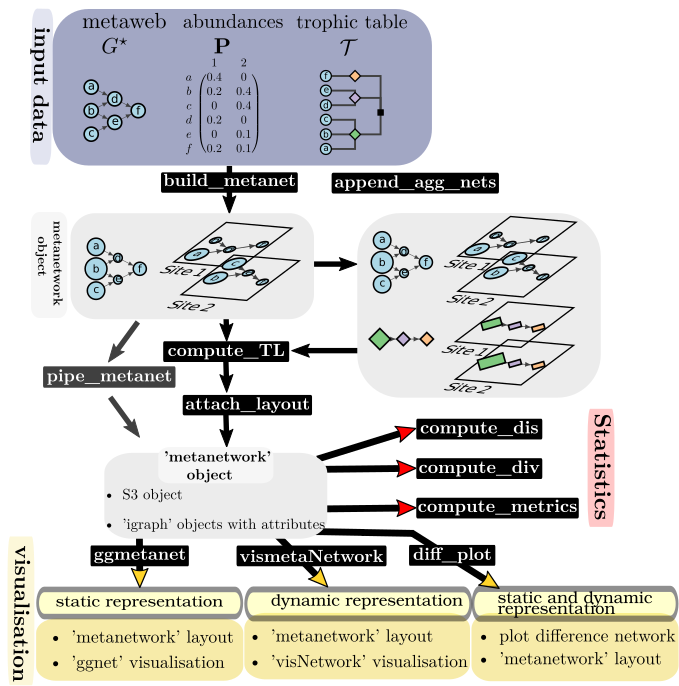
In ecological networks literature, metanetwork refers to a set of networks in space. In R package ‘metanetwork’, we stick to a widespread (however restrictive) case:
- a potential interaction network (the metaweb, can be built using expert knowledge)
- local abundance tables, local networks are then induced subgraph of the metaweb by local abundances
Additional information might be considered (and used in ‘metanetwork’) as:
- a trophic table indicating a hierarchy of nodes of the metaweb, in order to study the metanetwork at different aggregation levels
See vignettes for application of ‘metanetwork’ on several datasets.